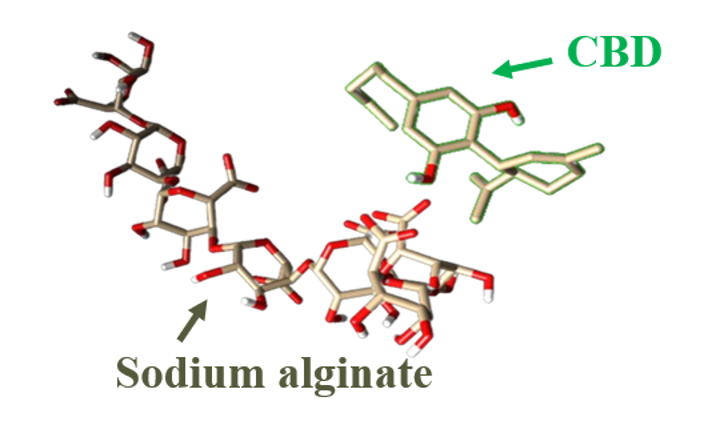
Molecular Dynamics Study to Analyze the Interactions between Sodium Alginate and CBD
DOI:
https://doi.org/10.29356/jmcs.v70i1.2440Keywords:
Molecular dynamics, CBD, alginate, encapsulationAbstract
Sodium alginate is a hydrophilic polysaccharide, widely recognized for its biocompatibility, making it suitable for applications in the human body. Its growing use as a matrix for encapsulating hydrophobic molecules and in controlled drug release highlights its research potential. Cannabinoids, on the other hand, are generally hydrophobic; cannabidiol (CBD) stands out for its therapeutic properties, attracting significant interest in recent years. In this study, molecular dynamics simulations were used to investigate the interactions between sodium alginate and CBD in water and a simulated CaCl₂ saline solution to assess their affinity and the potential of alginate as an encapsulation matrix. Both systems were evaluated under identical simulation conditions to observe interactions within the CBD-SA complexes. Molecular parameterization employed the OPLS-AA force field, with simulations running for 100 ns. The results revealed significant interactions in saline and aqueous environments, with differences suggesting the optimal physiological medium for CBD encapsulation. Overall, sodium alginate showed limitations in fully encapsulated CBD due to inconsistent interactions, although cases are highlighted where alginate combined with other compounds showed promising results.
Resumen. El alginato de sodio, es un polisacárido hidrofílico, ampliamente reconocido por su biocompatibilidad, lo que lo hace adecuado para aplicaciones en el cuerpo humano. Su creciente uso como matriz para encapsular moléculas hidrofóbicas y en la liberación controlada de fármacos destaca su potencial de investigación. Los cannabinoides, en cambio, son generalmente hidrofóbicos; el cannabidiol (CBD) sobresale por sus propiedades terapéuticas, atrayendo gran interés en los últimos años. En este estudio, se usaron simulaciones de dinámica molecular para investigar las interacciones entre el alginato de sodio y el CBD en agua y en una solución salina simulada de CaCl₂, con el fin de evaluar su afinidad y el potencial del alginato como matriz para la encapsulación. Ambos sistemas fueron evaluados bajo condiciones de simulación idénticas para observar interacciones en los complejos CBD-SA. La parametrización molecular utilizó el campo de fuerza OPLS-AA y las simulaciones se realizaron durante 100 ns. Los resultados revelaron interacciones significativas en entornos salino y acuoso, con diferencias que sugieren el medio fisiológico óptimo para la encapsulación de CBD. En general, el alginato de sodio mostró limitaciones para encapsular completamente el CBD debido a interacciones inconsistentes, aunque se destacan casos donde el alginato combinado con otros compuestos mostró resultados prometedores.
Downloads
References
1. Scuderi, C.; De Filippis, D.; Iuvone, T.; Blasio, A.; Steardo, A.; Esposito, G. Phytother. Res. 2009 23, 597–602. DOI: https://doi.org/10.1002/ptr.2625.
2. Seltzer, E. S.; Watters, A. K.; MacKenzie, D.; Granat, L. M.; Zhang, D. Cancers. 2020 12, 3203. DOI: https://doi.org/10.3390/cancers12113203.
3. Peng, J.; Fan, M.; An, C.; Ni, F.; Huang, W.; Luo, J. Basic Clin. Pharmacol. Toxicol. 2022 130, 439–456. DOI: https://doi.org/10.1111/bcpt.13710.
4. Francke, N. M.; Schneider, F.; Baumann, K.; Bunjes, H. Molecules. 2021 26, 1469. DOI: https://doi.org/10.3390/molecules26051469.
5. Grifoni, L.; Vanti, G.; Donato, R.; Sacco, C.; Bilia, A. R. Molecules. 2022 27. DOI: https://doi.org/10.3390/molecules27186070.
6. Wang, C.; Cui, B.; Sun, Y.; Wang, C.; Guo, M. LWT. 2022 162, 113466. DOI: https://doi.org/10.1016/j.lwt.2022.113466.
7. Noreen, S.; et al. Drug Deliv. Transl. Res. 2022 12, 2649–2666. DOI: https://doi.org/10.1007/s13346-022-01152-3.
8. Villate, A.; San Nicolas, M.; Olivares, M.; Aizpurua-Olaizola, O.; Usobiaga, A. Pharmaceutics. 2023 15, 859. DOI: https://doi.org/10.3390/pharmaceutics15030859.
9. Plazinski, W.; Plazinska, A. New J. Chem. 2011 35, 1607. DOI: https://doi.org/10.1039/C1NJ20273A.
10. Perić-Hassler, L.; Hünenberger, P. H. Mol. Simul. 2010 36, 778–795. DOI: https://doi.org/10.1080/08927021003752853.
11. da Silva, A. J.; dos Santos, E. S. Eur. Biophys. J. 2020 49, 571–589. DOI: https://doi.org/10.1007/s00249-020-01463-8.
12. Hollingsworth, S. A.; Dror, R. O. Neuron. 2018 99, 1129–1143. DOI: https://doi.org/10.1016/j.neuron.2018.08.011.
13. Fleten, K. G.; et al. Mar. Drugs. 2022 20, 744. DOI: https://doi.org/10.3390/md20120744.
14. Hecht, H.; Srebnik, S. Biomacromolecules. 2016 17, 2160–2167. DOI: https://doi.org/10.1021/acs.biomac.6b00378.
15. Xiang, Y.; Liu, Y.; Mi, B.; Leng, Y. Langmuir. 2014 30, 9098–9106. DOI: https://doi.org/10.1021/la501811d.
16. Hanwell, M. D.; Curtis, D. E.; Lonie, D. C.; Vandermeersch, T.; Zurek, E.; Hutchison, G. R. J. Cheminform. 2012 4, 17. DOI: https://doi.org/10.1186/1758-2946-4-17.
17. O’Boyle, N. M.; Banck, M.; James, C. A.; Morley, C.; Vandermeersch, T.; Hutchison, G. R. J. Cheminform. 2011 3, 33. DOI: https://doi.org/10.1186/1758-2946-3-33.
18. Dodda, L. S.; Cabeza de Vaca, I.; Tirado-Rives, J.; Jorgensen, W. L. Nucleic Acids Res. 2017 45, W331–W336. DOI: https://doi.org/10.1093/nar/gkx312.
19. Dodda, L. S.; Vilseck, J. Z.; Tirado-Rives, J.; Jorgensen, W. L. J. Phys. Chem. B 2017 121, 3864–3870. DOI: https://doi.org/10.1021/acs.jpcb.7b00272.
20. Jorgensen, W. L.; Tirado-Rives, J. Proc. Natl. Acad. Sci. U.S.A. 2005 102, 6665–6670. DOI: https://doi.org/10.1073/pnas.0408037102.
21. Kony, D.; Damm, W.; Stoll, S.; Van Gunsteren, W. F. J. Comput. Chem. 2002 23, 1416–1429. DOI: https://doi.org/10.1002/jcc.10139.
22. Damm, W.; Frontera, A.; Tirado-Rives, J.; Jorgensen, W. L. J. Comput. Chem. 1997 18, 1955–1970. DOI: https://doi.org/10.1002/(SICI)1096-987X(199712)18:16<1955::AID-JCC1>3.0.CO;2-L.
23. Eberhardt, J.; Santos-Martins, D.; Tillack, A. F.; Forli, S. J. Chem. Inf. Model. 2021 61, 3891–3898. DOI: https://doi.org/10.1021/acs.jcim.1c00203.
24. Trott, O.; Olson, A. J. J. Comput. Chem. 2010 31, 455–461. DOI: https://doi.org/10.1002/jcc.21334.
25. Abraham, M.; et al. GROMACS 2023.1 Manual. 2023.
26. Bussi, G.; Donadio, D.; Parrinello, M. J. Chem. Phys. 2007 126. DOI: https://doi.org/10.1063/1.2408420.
27. Hess, B.; Bekker, H.; Berendsen, H. J. C.; Fraaije, J. G. E. M. J. Comput. Chem. 1997 18, 1463–1472. DOI: https://doi.org/10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H.
28. Majimbi, M.; et al. PLoS One. 2021 16, e0243858. DOI: https://doi.org/10.1371/journal.pone.0243858.

Downloads
Published
Issue
Section
License
Copyright (c) 2026 Erik De La Rosa Montelongo, Lucero Rosales-Marines, Lorena Farías-Cepeda, Juan De La Peña-Zúñiga

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Authors who publish with this journal agree to the following terms:
- Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution License that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
- Authors are able to enter into separate, additional contractual arrangements for the non-exclusive distribution of the journal's published version of the work (e.g., post it to an institutional repository or publish it in a book), with an acknowledgement of its initial publication in this journal.